
Research
We investigate the molecular mechanisms that enable the human pathogen Mycobacterium tuberculosis to establish and maintain chronic infections and resist host defenses. M. tuberculosis has evolved to resist killing by macrophages, its primary host cells. It takes months to treat drug susceptible tuberculosis (TB) and years if the pathogen has acquired antibiotic resistance. We apply genetic approaches to identify the pathways that the pathogen requires for intracellular growth and survival. These pathways could be targeted to kill populations that escape current chemotherapy in people with active or latent M. tuberculosis infection. We also analyze host immune factors that are required to control M. tuberculosis and seek to identify new strategies to generate improved TB vaccines.
Figure 1
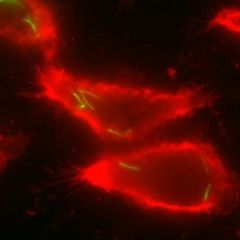
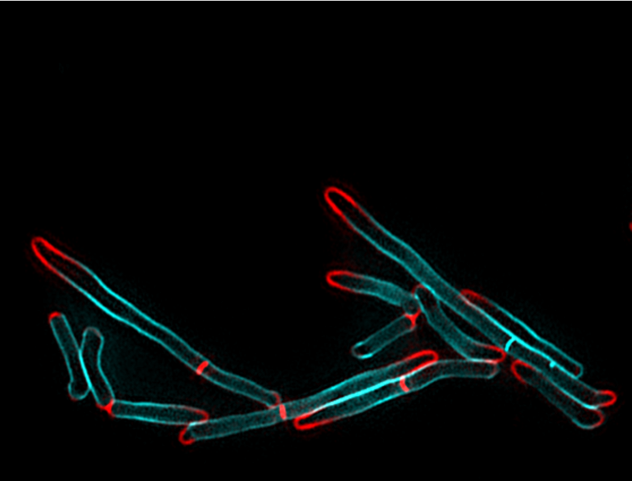
Figure 2
Current Projects:
- Mechanisms of mycobacterial intracellular growth and survival
- Latent M. tuberculosis infection
- Metabolic adaptation and nutrient acquisition during infection
- Drug target identification
- TB vaccines
Bio
Sabine Ehrt received her Ph.D. in Microbiology from the Friedrich-Alexander University of Erlangen-Nürnberg, Germany. As a postdoctoral fellow she started her work on Mycobacterium tuberculosis at Cornell University Medical College and then at the University of California at Berkeley. Dr. Ehrt joined the faculty in the Department of Microbiology and Immunology at Weill Cornell Medical College in 1999. She is currently Professor of Microbiology and Immunology and Co-Chair of the Immunology and Microbial Pathogenesis Graduate Program.
Distinctions:
- Outstanding Teaching and Mentoring Award (WCGS, 2020)
- Fellow, American Academy of Microbiology (2015)
- Excellence in Mentoring (WCM Postdoctoral Association, 2006)
- Career Scientist Award, Irma T. Hirschl Trust (2005)
